Arabinose operon is one of the regulatory systems found in the bacterial cell (E.coli), facilitating arabinose catalysis. L-arabinose operon and ARA-operon are the two alternative names of the arabinose operon. Ara-operon system provides energy to the cell by the breakdown of arabinose into xylulose 5-phosphate.
Arabinose is a 5-C sugar or aldopentose that provides energy in a carbon source to the bacterial cells. Xylulose 5-phosphate is a ketose sugar and serves as an intermediary product of the pentose phosphate pathway. The bacteria utilize this carbon source to perform various functions. In the arabinose operon, there is a group of genes that perform specific functions within the cell.
The structural genes (ara-bad genes) encodes for three enzymes that aid in the degradation of arabinose. Ribulokinase (ara-B), I-arabinose isomerase (ara-A) and I-ribulose 5-phosphate 4-epimerase (ara-D) are the enzymes encodes by the ara-bad structural genes. Ara-C regulates the arabinose operon. Ara O1 and O2 are the operator genes, and PBAD and PC are the ara-operon’s promoter genes.
The main purpose of arabinose operon is to break the complex arabinose molecule into xylulose 5-phosphate, which then enters the metabolic pathway (pentose phosphate pathway). You will get to know the definition, structural elements and regulation of the arabinose operon.
Content: Arabinose Operon
Definition of Arabinose Operon
Arabinose operon can define the system carrying the number of genes like a regulatory, promoter, operator, inducer, and structural genes for L-Arabinose’s breakdown into xylulose 5-phosphate. The structural genes ara-B, A and D carry out the conversion of arabinose into xylulose 5-phosphate. Ara-B, Ara-A and Ara-D transcribe the mRNA by encoding the kinase, isomerase and epimerase enzymes. The enzymes kinase, isomerase, and epimerase catalyze L-arabinose’s conversion to xylulose phosphate, which then enters the pentose phosphate pathway.
Structural Elements
The structure of arabinose operon is linear, and it consists of four specific genes along with the catabolic active site.
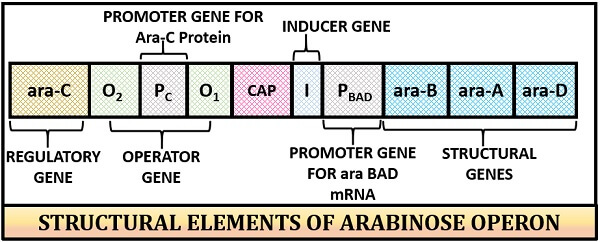
- Structural genes
- Inducer genes
- Catabolic active site
- Operator genes
- Promoter genes
- Regulatory gene
Structural Genes
There are three types of structural genes, namely ara-B, ara-A and ara-D. The main function of the structural genes involves the encoding of the metabolic enzymes. The metabolic enzymes (kinase, isomerase and epimerase) are synthesized by the ara-B, ara-A and ara-D, respectively. The metabolic enzymes breakdown the non-glucose molecule, i.e. arabinose to produce a multigenic or polycistronic mRNA.
Inducer Gene
Ara I acts as an inducer gene. The ara I gene induces the transcription. In arabinose operon, the inducer molecule is the arabinose that binds with the repressor protein to induce the transcription of the gene into mRNA. The arabinose operon works positively in the presence of inducer and works negatively in the absence of inducer.
Catabolic Active Site
In arabinose operon, an activator site is termed as CAP. A term CAP stands for catabolite activator protein, which activates the efficiency of transcription rate by promoting the effective binding of the RNA polymerase to the promoter region. When the availability of glucose is high with low arabinose, the ATP will not convert into cAMP.
If the availability of glucose is low with high arabinose content, then the ATP will convert into cAMP, and the whole complex will then bind to the CAP to activate the transcription of mRNA. The cyclic AMP plus CAP will form a complex which binds to the CAP region of the operon.
Operator Gene
There are two operator genes, namely O1 and O2 (operate araBAD mRNA synthesis). In positive regulation, when the amount of glucose is low, and the amount of arabinose is high, the repressor protein (Ara C) will get activated by the arabinose. It will promote the synthesis of araBAD mRNA. In negative regulation, when the amount of glucose is high, and the amount of arabinose is low, the repressor protein (Ara C) will not promote araBAD mRNA synthesis.
Promoter Gene
There are two promoter genes in the arabinose operon, namely PBAD and PC. PBAD is the promoter site of ara BAD structural genes, promoting the synthesis of ara BAD mRNA. PC is the promoter site of ara C regulatory gene, promoting the Ara C repressor protein synthesis. The Ara C repressor exists in active P1 state (in the absence of inducer), while in inactive P2 state (in the presence of inducer).
Regulatory gene
The ara-C is the only regulatory gene in the arabinose operon. The ara-C gene encodes the Ara C protein that acts as a repressor. The Ara C protein regulates the arabinose operon both positively and negatively. When the Ara C protein binds with the operator, it will repress the synthesis of araBAD mRNA.
It will not promote the binding of RNA polymerase to the promoter region. When the Ara C repressor protein binds with the inducer, i.e. arabinose, the complex becomes activated. This activation of Ara C with the arabinose will promote the attachment of RNA polymerase to the promoter region and leads to the synthesis of araBAD mRNA.
Regulation of Arabinose Operon
The arabinose operon is regulated both positively and negatively, like a Lac-operon model.
Positive Regulation
Here, a term positive indicates that the synthesis of mRNA will occur. Therefore, the ara BAD mRNA is formed in positive regulation. Arabinose operon is positively regulated by the two conditions that are explained below:
Case-I (when both inducer and repressor protein is absent):
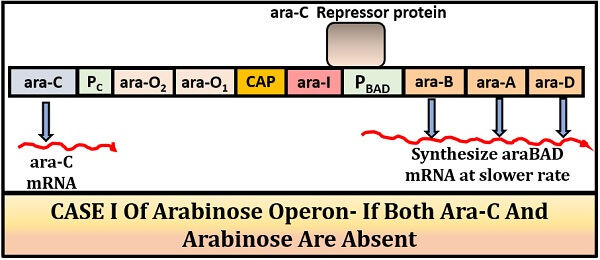
There will be no repression of the arabinose operon in the absence of both an inducer and a repressor protein. In this condition, the RNA polymerase will bind with the specific promoter region and transcribe the ara-BAD genes to form mRNA. But in this case, the rate of transcribing mRNA is much slower.
Case-II (when both inducer and repressor are present):
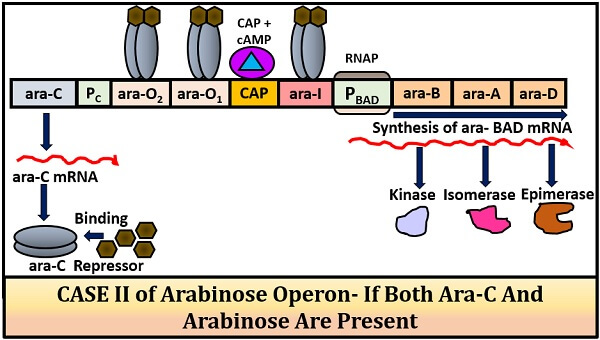
The inducer (arabinose) will bind with the repressor protein to regulate the mRNA transcription. The Ara-C protein plus arabinose forms a complex that will not allow the loop formation. The arabinose binds with the Ara-C dimer and changes its structural configuration. This change in structural configuration will allow the RNA polymerase to transcribe the ara-BAD genes to form mRNA. The mRNA will further translate into proteins.
Negative Regulation
Here, a term negative indicates that the mRNA transcription will not occur. To understand the process in a simple way, let us take condition III to compare it with the before two conditions.
Case-III (when only repressor protein is present):
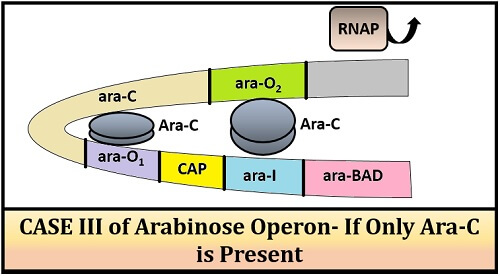
The main role of arabinose operon involves the breakdown of arabinose. In the absence of arabinose, there will be no transcription and translation of the DNA molecule. Arabinose acts as an inducer, which binds and inactivates the repressor protein ARA-C. In the absence of inducer, the repressor protein will be produced by the ara-C gene. ARA-C repressor protein forms a dimer with the operator and the inducer gene by forming a loop. The loop formation will not allow the RNA polymerase to transcribe the ara-BAD genes to form mRNA.
Conclusion
Therefore, ara-operon can be both positively and negatively regulated. In simple words, we can say that the arabinose operon system can be controlled by both activation and repression. The repression of the operon is through the repressor protein, i.e. Ara-C protein. The activation of the operon is through the inducer. So, we can conclude that the arabinose operon is switched off in the presence of repressor and switched on in the presence of inducer.