Rho protein or Rho factor is a hexamer complex, which is ATP-dependent. Rho gene synthesizes rho protein to terminate the transcription process in prokaryotes. It is a protein possessing a single polypeptide chain with 419 amino acid residues.
It functions as an RNA-binding protein, which primarily recognizes and binds to the rut site of the newly synthesized or nascent RNA. Rho utilization site (Ruts) serves as an activator and loader of rho protein.
After binding, the CTD of rho factor facilitates ATP hydrolysis and hydrolysis of other nucleoside triphosphates. Subsequently, Rho runs along the length of nascent RNA and ultimately reaches the core enzyme of transcription (RNA polymerase) by utilizing its ATP hydrolysis activity as a source of energy.
Then, it dislodges the RNA transcript from RNA polymerase and the DNA template. In this post, we will study the definition, rho-dependent termination and rho-independent termination in the transcription of the prokaryotic genome. The structure and significant functions of the rho protein have also been explained.
Content: Rho Protein
Definition of Rho Protein
Rho protein (Rho factor) refers to a transcription termination protein found in the majority of prokaryotes. It exists as a well-conserved protein that terminates the multistep transcription process.
Rho factor dislodges the elongated the transcribed mRNA and RNA-polymerase from the DNA template strand at specific sites. Prokaryotes have two sets of transcriptional terminators, namely intrinsic and rho dependent.
Rho protein exists as a hexamer and RNA dependent ATPase. Some bacteria like Bacilli, Mollicutes, Clostridia lack the rho factor and undergo intrinsic termination (without a rho protein).
Conversely, the members belonging to the Enterobacteriaceae family possess rho protein and undergo rho-dependent transcription termination.
Rho Dependent Termination
It is mediated by the rho factor. Rho gene encodes Rho proteins. Rho protein is a hexamer. A termination site is located near the promoter region of the DNA template. Although, the termination site does not cause termination of mRNA transcription.
Rho protein facilitates termination by binding with the rut-site on the sRNA or nascent RNA. Then, the Rho translocates along with the RNA and reaches the RNA polymerase enzyme through ATP hydrolysis. The single stranded RNA winds around the Rho hexamer.
This whole complex precedes to the core enzyme of polymerase. Nus A and NusG proteins aids in the association of the wrapped rho protein and the RNAP by utilizing energy from ATP hydrolysis.
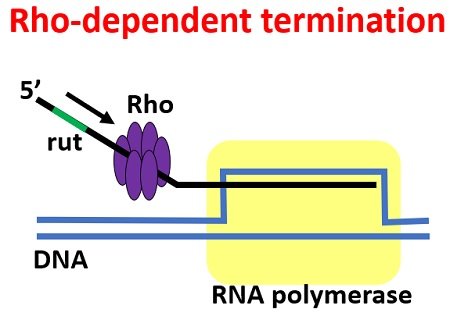
The twisting of nascent RNA around the rho protein continues, which causes the weakening of the non-covalent bonds holding the newly synthesized strand of the RNA with the DNA template strand.
Eventually, the rho protein through its helicase activity results in the dissociation of the newly formed RNA and the RNA polymerase enzyme from the DNA template. Therefore, the rho-dependent transcription termination occurs in this way.
Rho Independent Termination
It is controlled by the specific sequences (C–G rich) in the newly formed RNA transcript. The RNA polymerase reads and extends the poly-A sequence on the DNA template strand and synthesizes an RNA transcript accordingly.
The transcribed RNA transcript (mRNA) possesses some palindromic sequences C and G sequences and poly-U sequences. The complementary C–G nucleotides in the mRNA transcript complementarily pair with each other.
It results in the folding back of the mRNA transcript on itself, and a sturdy stem and loop or hairpin structure forms due to strongly associated C and G nucleotides.
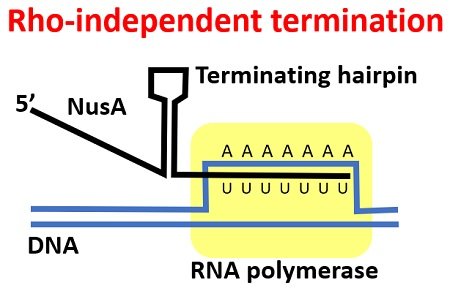
The hairpin structure in the newly synthesized RNA ceases the RNA polymerase reading mechanism and its further movement on the DNA template.
The complementary U–A nucleotides pairing between the mRNA transcript and the DNA template forms a weak site. As a result of weak interaction, the new mRNA transcript falls.
Rho Protein Structure
Rho has six laterally arranged identical subunits. Each subunit has ATP-binding sites located (at the interface of the C-terminal), N and C-terminal domains. ATP binding sites are composed of Rec-A like proteins that utilize NTP.
Rho subunits appear peanut-shaped. The N-terminal domain has alpha subdomain (alpha helical bundle) and beta subdomain (beta-barrel). The C-terminal domain has parallel beta-sheets between several alpha-helices.
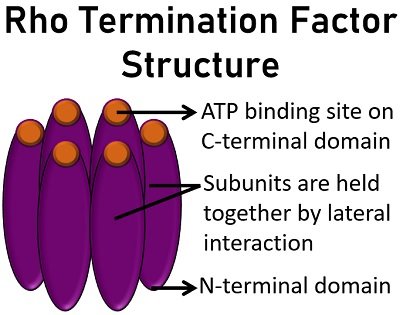
In addition, C-terminal has three key motifs, namely P, R and Q loops. P-loop provides a site for ATP binding and ATP hydrolysis and comprises 179 to 183 amino acids. Oppositely, R loop (322 amino acids) and Q loop (278 amino acids) offers a location for the binding of secondary RNA.
Six subunits are catalytically competent and associated by lateral interactions. The subunits hydrolyze ATP sequentially. Rho binds to the cytosine-rich rut site.
Function
The primary function of rho factor is to terminate the gene expression in prokaryotes. Rho factors function as the DNA/RNA helicases that dissociate the newly synthesized mRNA transcript, thereby terminating transcription.
Rho binds to the rut site on the nascent mRNA once translation encounters nutrient scarcity or nonsense mutations. It also regulates the action of sRNA (small non coding bacterial RNA). The newly formed single stranded RNA have the potential to form secondary structures (hairloops).
Rho removes the stem and loop structures in the newly synthesized RNA through its helicase activity, thereby promoting RNA remodelling.